Iodine »
PDB 1gjd-1lij »
1gth »
Iodine in PDB 1gth: Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil
Enzymatic activity of Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil
All present enzymatic activity of Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil:
1.3.1.2;
1.3.1.2;
Protein crystallography data
The structure of Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil, PDB code: 1gth
was solved by
D.Dobritzsch,
S.Ricagno,
G.Schneider,
K.D.Schnackerz,
Y.Lindqvist,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 25.00 / 2.25 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 82.210, 159.690, 167.590, 90.00, 97.11, 90.00 |
| R / Rfree (%) | 17.7 / 20.9 |
Other elements in 1gth:
The structure of Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil also contains other interesting chemical elements:
| Iron | (Fe) | 64 atoms |
Iodine Binding Sites:
The binding sites of Iodine atom in the Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil
(pdb code 1gth). This binding sites where shown within
5.0 Angstroms radius around Iodine atom.
In total 2 binding sites of Iodine where determined in the Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil, PDB code: 1gth:
Jump to Iodine binding site number: 1; 2;
In total 2 binding sites of Iodine where determined in the Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil, PDB code: 1gth:
Jump to Iodine binding site number: 1; 2;
Iodine binding site 1 out of 2 in 1gth
Go back to
Iodine binding site 1 out
of 2 in the Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil
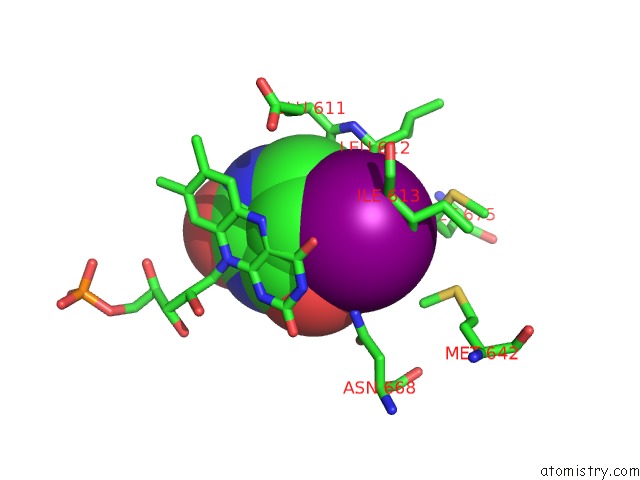
Mono view
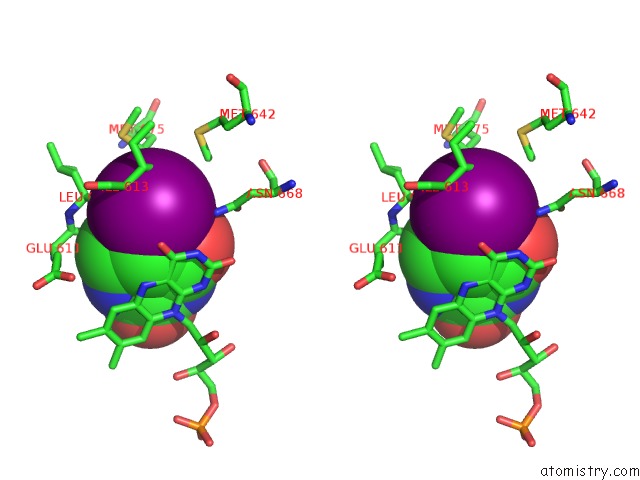
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 1 of Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil within 5.0Å range:
|
Iodine binding site 2 out of 2 in 1gth
Go back to
Iodine binding site 2 out
of 2 in the Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil
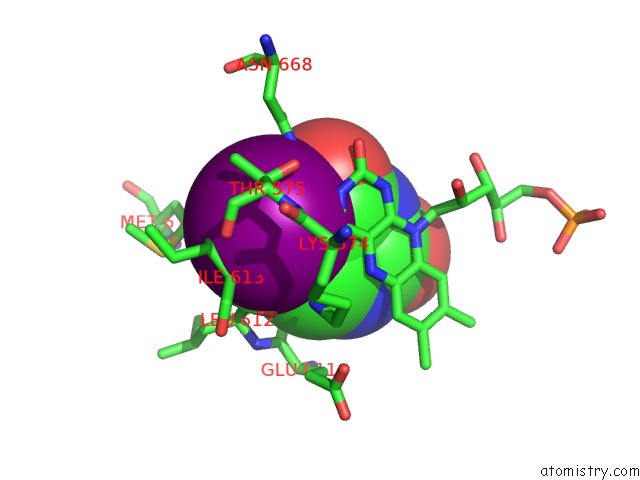
Mono view
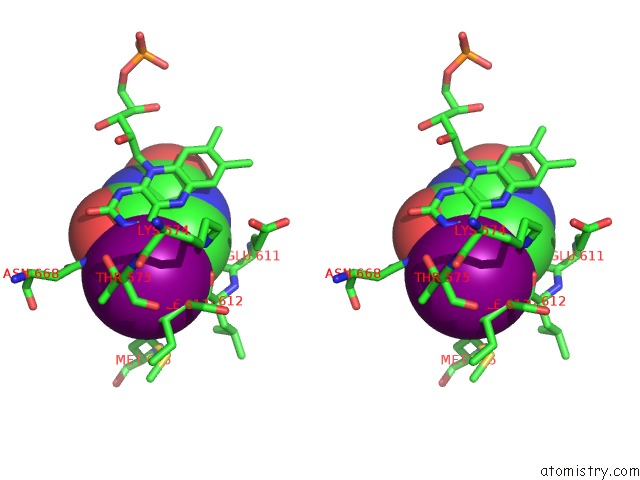
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 2 of Dihydropyrimidine Dehydrogenase (Dpd) From Pig, Ternary Complex with Nadph and 5-Iodouracil within 5.0Å range:
|
Reference:
D.Dobritzsch,
S.Ricagno,
G.Schneider,
K.D.Schnackerz,
Y.Lindqvist.
Crystal Structure of the Productive Ternary Complex of Dihydropyrimidine Dehydrogenase with Nadph and 5-Iodouracil. Implications For Mechanism of Inhibition and Electron Transfer. J. Biol. Chem. V. 277 13155 2002.
ISSN: ISSN 0021-9258
PubMed: 11796730
DOI: 10.1074/JBC.M111877200
Page generated: Fri Aug 8 11:48:40 2025
ISSN: ISSN 0021-9258
PubMed: 11796730
DOI: 10.1074/JBC.M111877200
Last articles
I in 3RTNI in 3RTM
I in 3RTH
I in 3RSX
I in 3RKE
I in 3RSV
I in 3PFD
I in 3REW
I in 3RIA
I in 3R5O