Iodine »
PDB 442d-4dch »
4cdw »
Iodine in PDB 4cdw: Crystal Structure of Human Enterovirus 71 in Complex with the Uncoating Inhibitor GPP4
Protein crystallography data
The structure of Crystal Structure of Human Enterovirus 71 in Complex with the Uncoating Inhibitor GPP4, PDB code: 4cdw
was solved by
L.De Colibus,
X.Wang,
J.A.B.Spyrou,
J.Kelly,
J.Ren,
J.Grimes,
G.Puerstinger,
N.Stonehouse,
T.S.Walter,
Z.Hu,
J.Wang,
X.Li,
W.Peng,
D.Rowlands,
E.E.Fry,
Z.Rao,
D.I.Stuart,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 49.63 / 2.80 |
| Space group | I 2 3 |
| Cell size a, b, c (Å), α, β, γ (°) | 599.660, 599.660, 599.660, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 28.3 / 28.5 |
Other elements in 4cdw:
The structure of Crystal Structure of Human Enterovirus 71 in Complex with the Uncoating Inhibitor GPP4 also contains other interesting chemical elements:
| Sodium | (Na) | 2 atoms |
Iodine Binding Sites:
The binding sites of Iodine atom in the Crystal Structure of Human Enterovirus 71 in Complex with the Uncoating Inhibitor GPP4
(pdb code 4cdw). This binding sites where shown within
5.0 Angstroms radius around Iodine atom.
In total only one binding site of Iodine was determined in the Crystal Structure of Human Enterovirus 71 in Complex with the Uncoating Inhibitor GPP4, PDB code: 4cdw:
In total only one binding site of Iodine was determined in the Crystal Structure of Human Enterovirus 71 in Complex with the Uncoating Inhibitor GPP4, PDB code: 4cdw:
Iodine binding site 1 out of 1 in 4cdw
Go back to
Iodine binding site 1 out
of 1 in the Crystal Structure of Human Enterovirus 71 in Complex with the Uncoating Inhibitor GPP4

Mono view
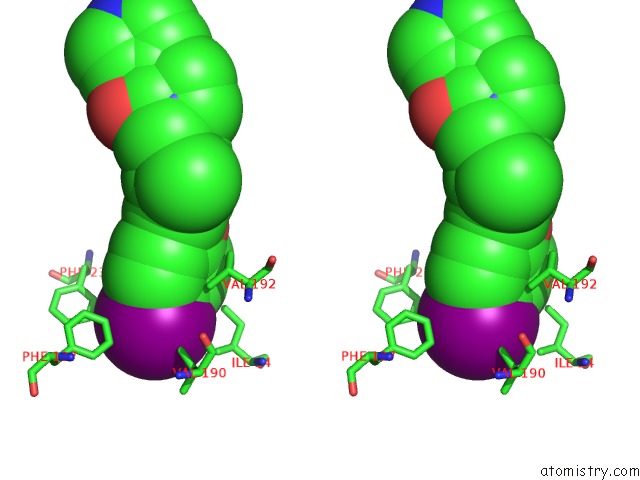
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 1 of Crystal Structure of Human Enterovirus 71 in Complex with the Uncoating Inhibitor GPP4 within 5.0Å range:
|
Reference:
L.De Colibus,
X.Wang,
J.A.B.Spyrou,
J.Kelly,
J.Ren,
J.Grimes,
G.Puerstinger,
N.Stonehouse,
T.S.Walter,
Z.Hu,
J.Wang,
X.Li,
W.Peng,
D.J.Rowlands,
E.E.Fry,
Z.Rao,
D.I.Stuart.
More-Powerful Virus Inhibitors From Structure-Based Analysis of HEV71 Capsid-Binding Molecules Nat.Struct.Mol.Biol. V. 21 282 2014.
ISSN: ISSN 1545-9993
PubMed: 24509833
DOI: 10.1038/NSMB.2769
Page generated: Fri Aug 8 16:52:46 2025
ISSN: ISSN 1545-9993
PubMed: 24509833
DOI: 10.1038/NSMB.2769
Last articles
Mg in 4DPGMg in 4DQP
Mg in 4DQQ
Mg in 4DPM
Mg in 4DPV
Mg in 4DQI
Mg in 4DOB
Mg in 4DOC
Mg in 4DMZ
Mg in 4DOA