Iodine »
PDB 4dgh-4hjh »
4e91 »
Iodine in PDB 4e91: Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3
Protein crystallography data
The structure of Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3, PDB code: 4e91
was solved by
C.T.Lemke,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 36.61 / 1.70 |
| Space group | P 32 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 65.783, 65.783, 143.323, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 22 / 24.3 |
Other elements in 4e91:
The structure of Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3 also contains other interesting chemical elements:
| Fluorine | (F) | 3 atoms |
| Chlorine | (Cl) | 2 atoms |
Iodine Binding Sites:
The binding sites of Iodine atom in the Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3
(pdb code 4e91). This binding sites where shown within
5.0 Angstroms radius around Iodine atom.
In total 4 binding sites of Iodine where determined in the Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3, PDB code: 4e91:
Jump to Iodine binding site number: 1; 2; 3; 4;
In total 4 binding sites of Iodine where determined in the Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3, PDB code: 4e91:
Jump to Iodine binding site number: 1; 2; 3; 4;
Iodine binding site 1 out of 4 in 4e91
Go back to
Iodine binding site 1 out
of 4 in the Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3
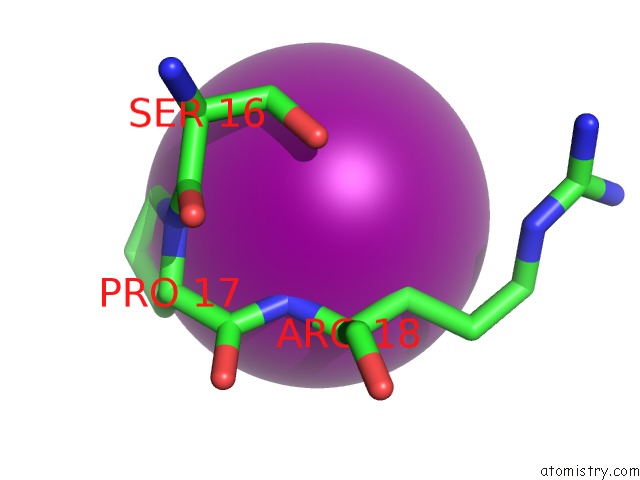
Mono view
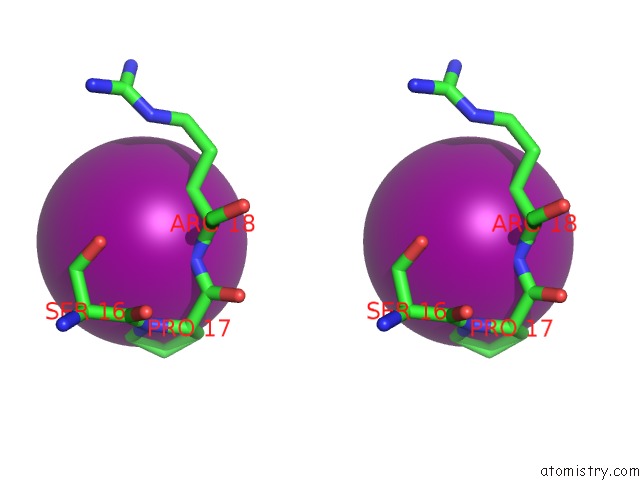
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 1 of Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3 within 5.0Å range:
|
Iodine binding site 2 out of 4 in 4e91
Go back to
Iodine binding site 2 out
of 4 in the Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3
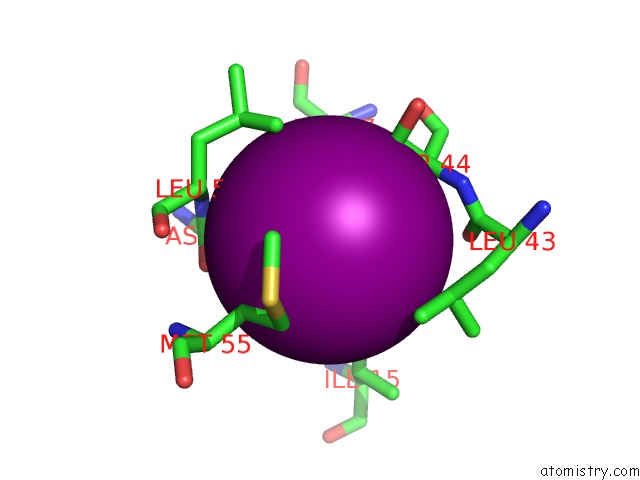
Mono view
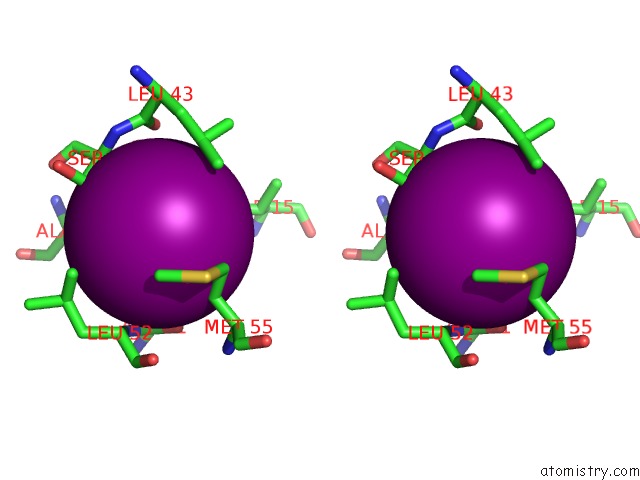
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 2 of Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3 within 5.0Å range:
|
Iodine binding site 3 out of 4 in 4e91
Go back to
Iodine binding site 3 out
of 4 in the Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3
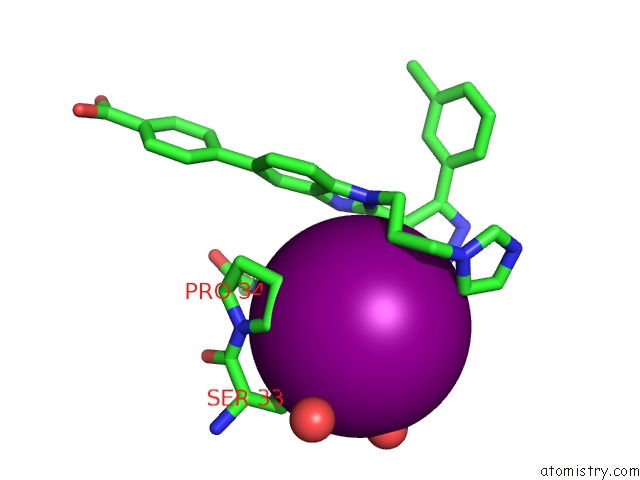
Mono view
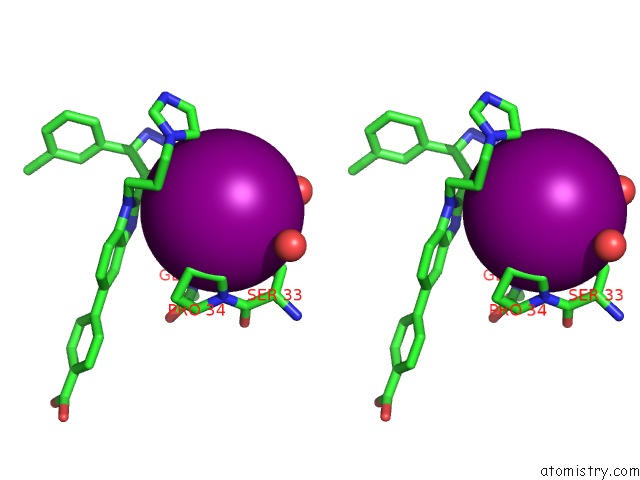
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 3 of Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3 within 5.0Å range:
|
Iodine binding site 4 out of 4 in 4e91
Go back to
Iodine binding site 4 out
of 4 in the Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3
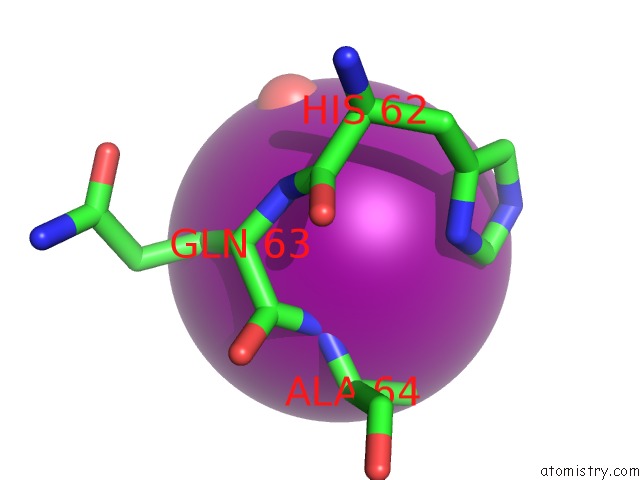
Mono view
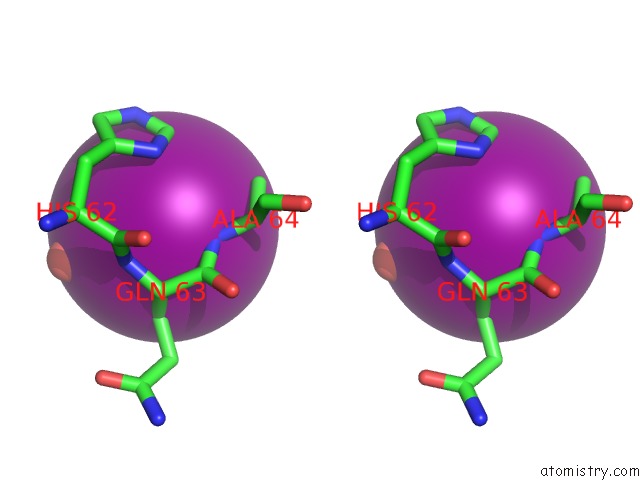
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 4 of Crystal Structure of the N-Terminal Domain of Hiv-1 Capsid in Complex with Inhibitor BD3 within 5.0Å range:
|
Reference:
C.T.Lemke,
S.Titolo,
U.Von Schwedler,
N.Goudreau,
J.F.Mercier,
E.Wardrop,
A.M.Faucher,
R.Coulombe,
S.S.Banik,
L.Fader,
A.Gagnon,
S.H.Kawai,
J.Rancourt,
M.Tremblay,
C.Yoakim,
B.Simoneau,
J.Archambault,
W.I.Sundquist,
S.W.Mason.
Distinct Effects of Two Hiv-1 Capsid Assembly Inhibitor Families That Bind the Same Site Within the N-Terminal Domain of the Viral Ca Protein. J.Virol. V. 86 6643 2012.
ISSN: ISSN 0022-538X
PubMed: 22496222
DOI: 10.1128/JVI.00493-12
Page generated: Sun Aug 11 17:36:31 2024
ISSN: ISSN 0022-538X
PubMed: 22496222
DOI: 10.1128/JVI.00493-12
Last articles
Ca in 5S5ECa in 5S5C
Ca in 5S5B
Ca in 5S5A
Ca in 5S59
Ca in 5S58
Ca in 5S57
Ca in 5S56
Ca in 5S55
Ca in 5S54