Iodine »
PDB 4dgh-4hjh »
4euu »
Iodine in PDB 4euu: Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172
Enzymatic activity of Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172
All present enzymatic activity of Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172:
2.7.11.1;
2.7.11.1;
Protein crystallography data
The structure of Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172, PDB code: 4euu
was solved by
X.Ma,
E.Helgason,
Q.T.Phung,
C.L.Quan,
R.S.Iyer,
M.W.Lee,
K.K.Bowman,
M.A.Starovasnik,
E.C.Dueber,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 50.00 / 1.80 |
| Space group | P 31 |
| Cell size a, b, c (Å), α, β, γ (°) | 76.095, 76.095, 130.945, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 16.9 / 19.4 |
Iodine Binding Sites:
The binding sites of Iodine atom in the Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172
(pdb code 4euu). This binding sites where shown within
5.0 Angstroms radius around Iodine atom.
In total 4 binding sites of Iodine where determined in the Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172, PDB code: 4euu:
Jump to Iodine binding site number: 1; 2; 3; 4;
In total 4 binding sites of Iodine where determined in the Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172, PDB code: 4euu:
Jump to Iodine binding site number: 1; 2; 3; 4;
Iodine binding site 1 out of 4 in 4euu
Go back to
Iodine binding site 1 out
of 4 in the Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172
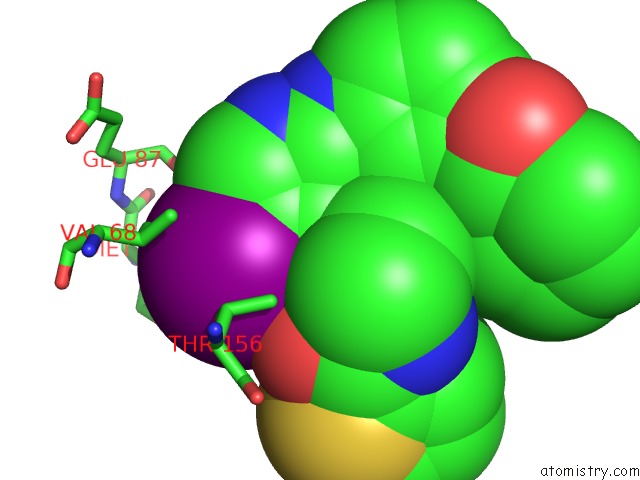
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 1 of Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172 within 5.0Å range:
|
Iodine binding site 2 out of 4 in 4euu
Go back to
Iodine binding site 2 out
of 4 in the Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172
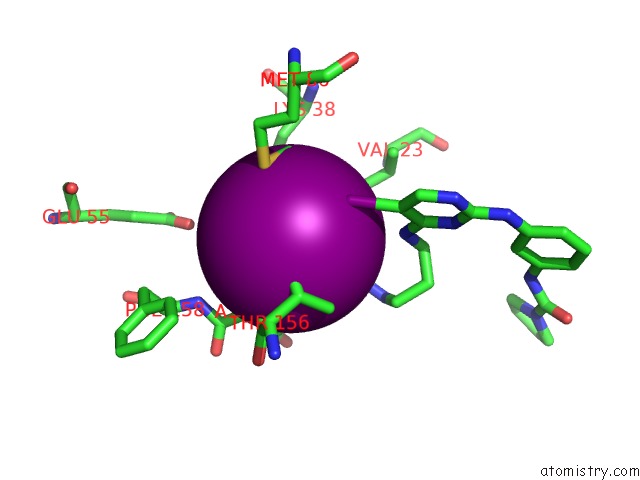
Mono view
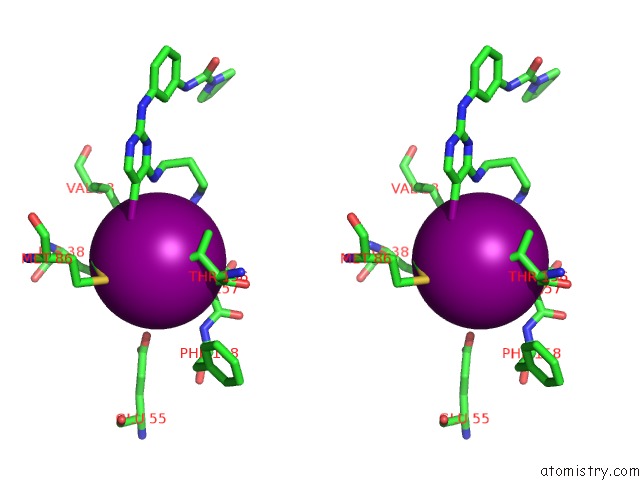
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 2 of Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172 within 5.0Å range:
|
Iodine binding site 3 out of 4 in 4euu
Go back to
Iodine binding site 3 out
of 4 in the Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172
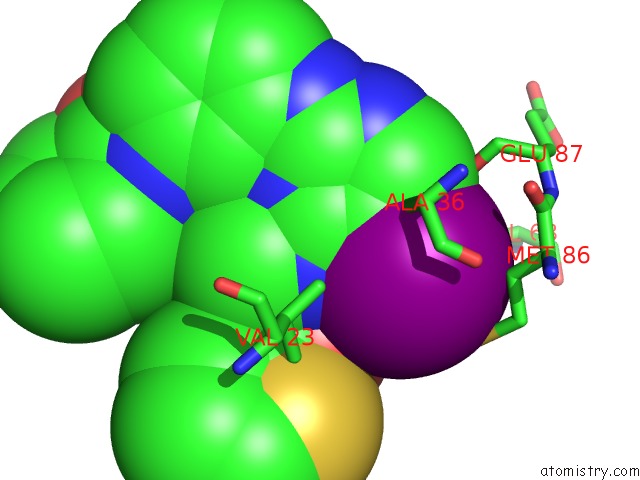
Mono view
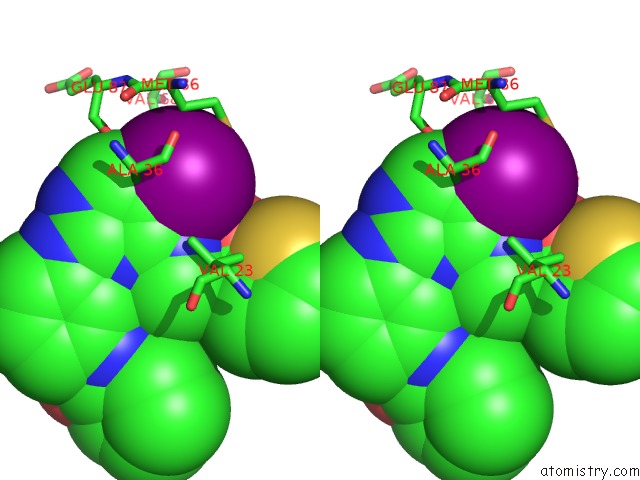
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 3 of Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172 within 5.0Å range:
|
Iodine binding site 4 out of 4 in 4euu
Go back to
Iodine binding site 4 out
of 4 in the Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 4 of Structure of Bx-795 Complexed with Human TBK1 Kinase Domain Phosphorylated on SER172 within 5.0Å range:
|
Reference:
X.Ma,
E.Helgason,
Q.T.Phung,
C.L.Quan,
R.S.Iyer,
M.W.Lee,
K.K.Bowman,
M.A.Starovasnik,
E.C.Dueber.
Molecular Basis of Tank-Binding Kinase 1 Activation By Transautophosphorylation. Proc.Natl.Acad.Sci.Usa V. 109 9378 2012.
ISSN: ISSN 0027-8424
PubMed: 22619329
DOI: 10.1073/PNAS.1121552109
Page generated: Fri Aug 8 17:01:26 2025
ISSN: ISSN 0027-8424
PubMed: 22619329
DOI: 10.1073/PNAS.1121552109
Last articles
K in 3GKOK in 3GVF
K in 3GJK
K in 3GE2
K in 3GAI
K in 3G71
K in 3GAJ
K in 3G6E
K in 3GAH
K in 3FWP