Iodine »
PDB 4n6d-4p4z »
4n6d »
Iodine in PDB 4n6d: Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site
Protein crystallography data
The structure of Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site, PDB code: 4n6d
was solved by
M.W.Vetting,
N.F.Al Obaidi,
M.Stead,
J.D.Attonito,
A.Scott Glenn,
S.Chowdhury,
B.Evans,
B.Hillerich,
J.Love,
R.D.Seidel,
H.J.Imker,
J.A.Gerlt,
S.C.Almo,
Enzyme Function Initiative (Efi),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 21.98 / 1.70 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 44.703, 68.249, 108.687, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 14.5 / 18.4 |
Other elements in 4n6d:
The structure of Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site also contains other interesting chemical elements:
| Chlorine | (Cl) | 1 atom |
Iodine Binding Sites:
The binding sites of Iodine atom in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site
(pdb code 4n6d). This binding sites where shown within
5.0 Angstroms radius around Iodine atom.
In total 6 binding sites of Iodine where determined in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site, PDB code: 4n6d:
Jump to Iodine binding site number: 1; 2; 3; 4; 5; 6;
In total 6 binding sites of Iodine where determined in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site, PDB code: 4n6d:
Jump to Iodine binding site number: 1; 2; 3; 4; 5; 6;
Iodine binding site 1 out of 6 in 4n6d
Go back to
Iodine binding site 1 out
of 6 in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site
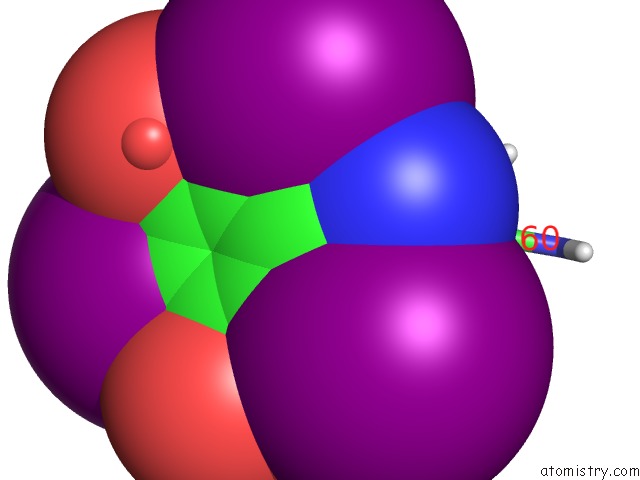
Mono view
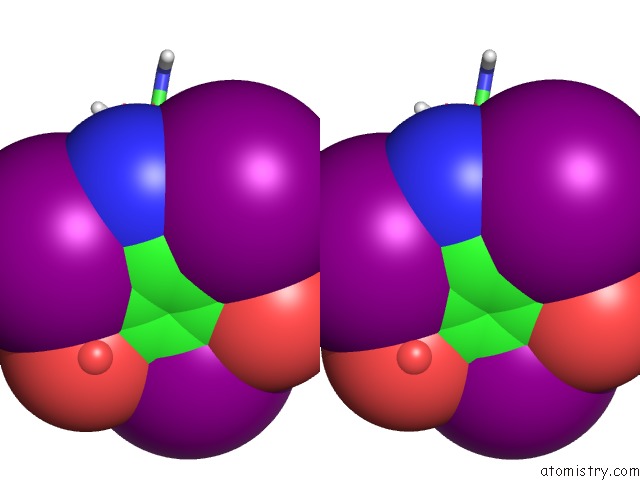
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 1 of Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site within 5.0Å range:
|
Iodine binding site 2 out of 6 in 4n6d
Go back to
Iodine binding site 2 out
of 6 in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site
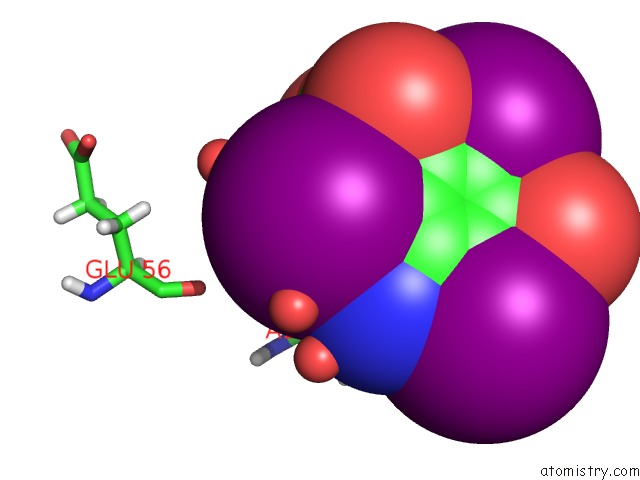
Mono view
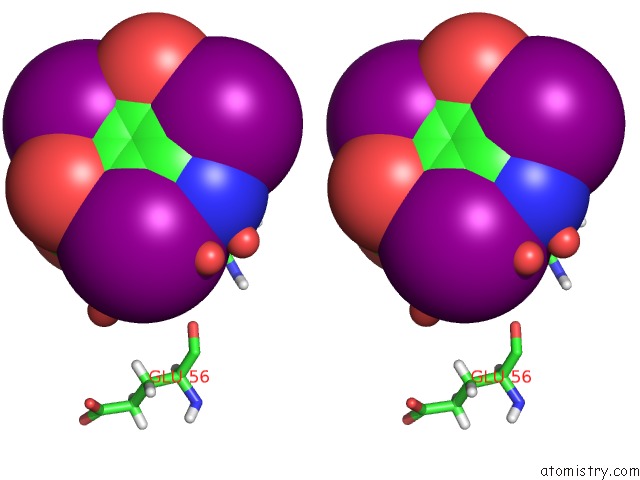
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 2 of Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site within 5.0Å range:
|
Iodine binding site 3 out of 6 in 4n6d
Go back to
Iodine binding site 3 out
of 6 in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site
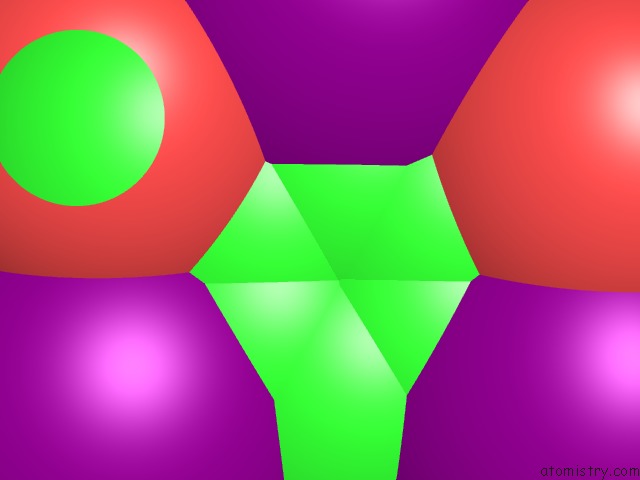

Mono view
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 3 of Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site within 5.0Å range:
|
Iodine binding site 4 out of 6 in 4n6d
Go back to
Iodine binding site 4 out
of 6 in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site
Mono view
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 4 of Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site within 5.0Å range:
|
Iodine binding site 5 out of 6 in 4n6d
Go back to
Iodine binding site 5 out
of 6 in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site
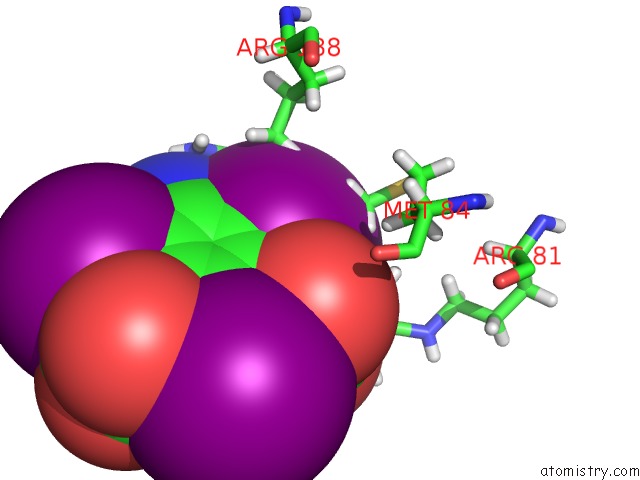
Mono view
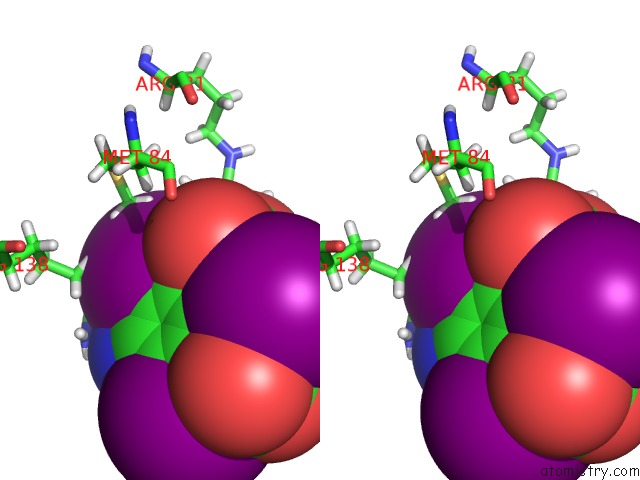
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 5 of Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site within 5.0Å range:
|
Iodine binding site 6 out of 6 in 4n6d
Go back to
Iodine binding site 6 out
of 6 in the Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site
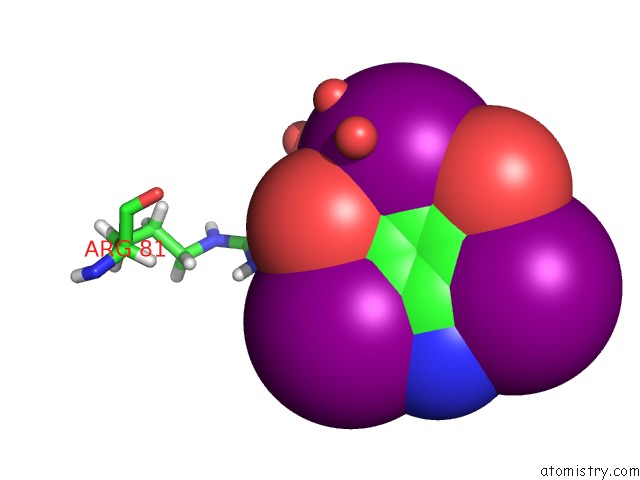
Mono view
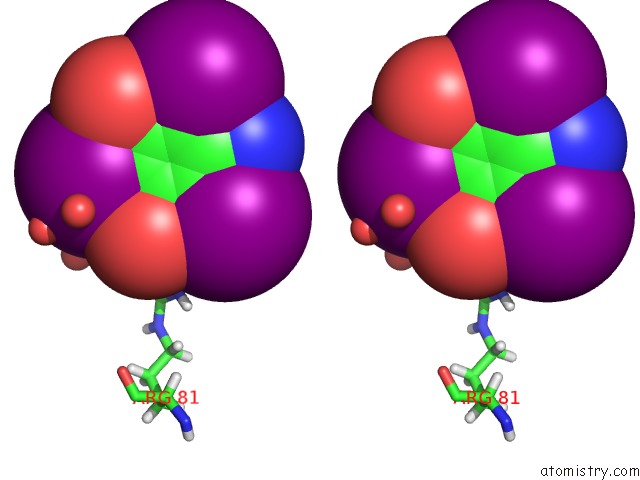
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 6 of Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site within 5.0Å range:
|
Reference:
M.W.Vetting,
N.F.Al Obaidi,
M.Stead,
J.D.Attonito,
A.Scott Glenn,
S.Chowdhury,
B.Evans,
B.Hillerich,
J.Love,
R.D.Seidel,
H.J.Imker,
J.A.Gerlt,
S.C.Almo.
Crystal Structure of A Trap Periplasmic Solute Binding Protein From Desulfovibrio Salexigens DSM2638 (DESAL_3247), Target Efi-510112, Phased with I3C, Open Complex, C-Terminus of Symmetry Mate Bound in Ligand Binding Site To Be Published.
Page generated: Sun Aug 11 18:53:24 2024
Last articles
Zn in 9J0NZn in 9J0O
Zn in 9J0P
Zn in 9FJX
Zn in 9EKB
Zn in 9C0F
Zn in 9CAH
Zn in 9CH0
Zn in 9CH3
Zn in 9CH1