Iodine »
PDB 7px1-7se4 »
7qur »
Iodine in PDB 7qur: Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
Iodine Binding Sites:
The binding sites of Iodine atom in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
(pdb code 7qur). This binding sites where shown within
5.0 Angstroms radius around Iodine atom.
In total 9 binding sites of Iodine where determined in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry, PDB code: 7qur:
Jump to Iodine binding site number: 1; 2; 3; 4; 5; 6; 7; 8; 9;
In total 9 binding sites of Iodine where determined in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry, PDB code: 7qur:
Jump to Iodine binding site number: 1; 2; 3; 4; 5; 6; 7; 8; 9;
Iodine binding site 1 out of 9 in 7qur
Go back to
Iodine binding site 1 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 1 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Iodine binding site 2 out of 9 in 7qur
Go back to
Iodine binding site 2 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
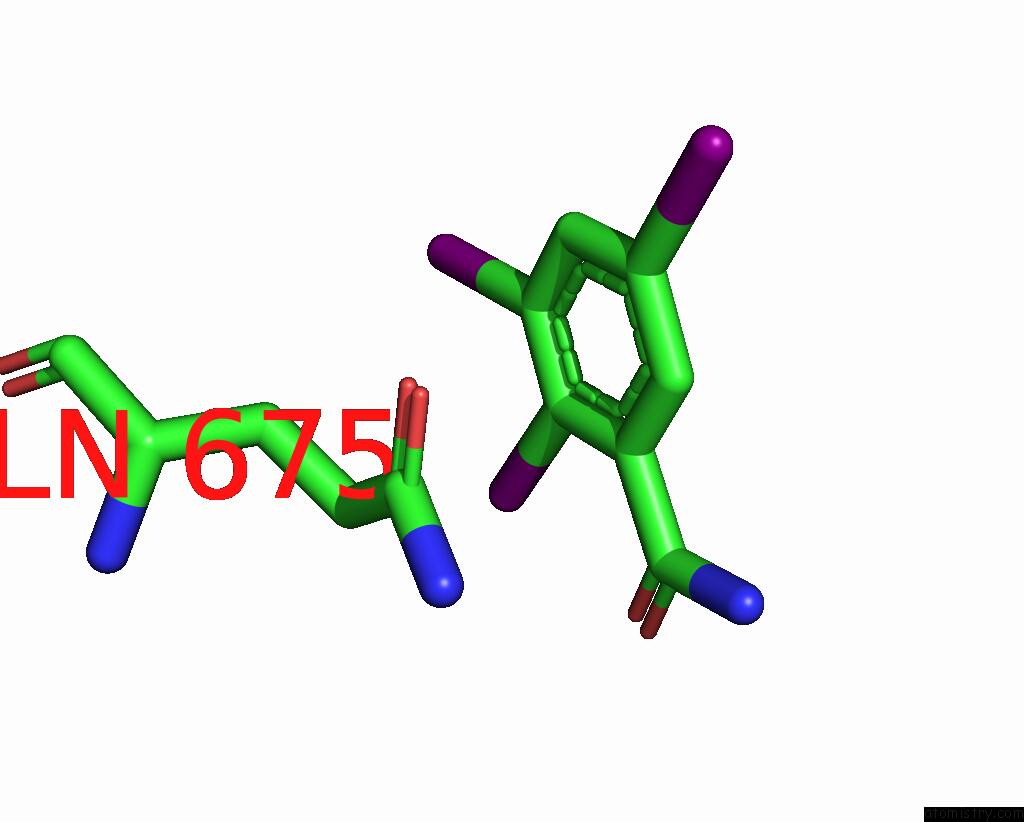
Mono view
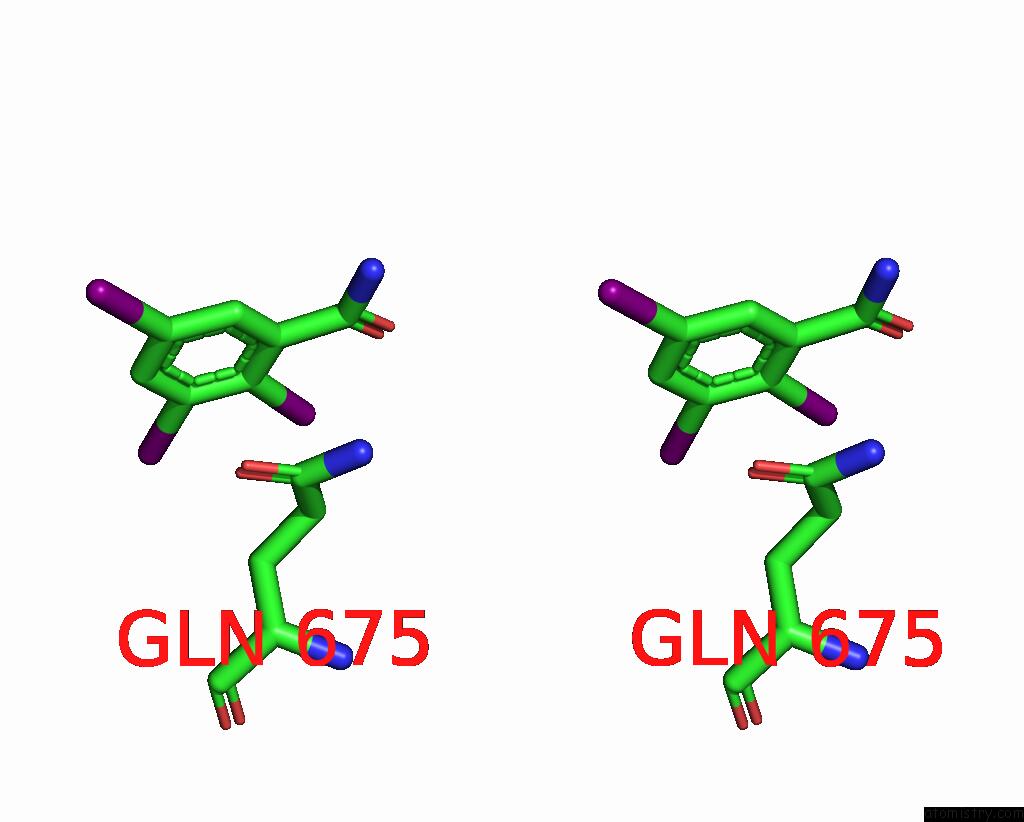
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 2 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Iodine binding site 3 out of 9 in 7qur
Go back to
Iodine binding site 3 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
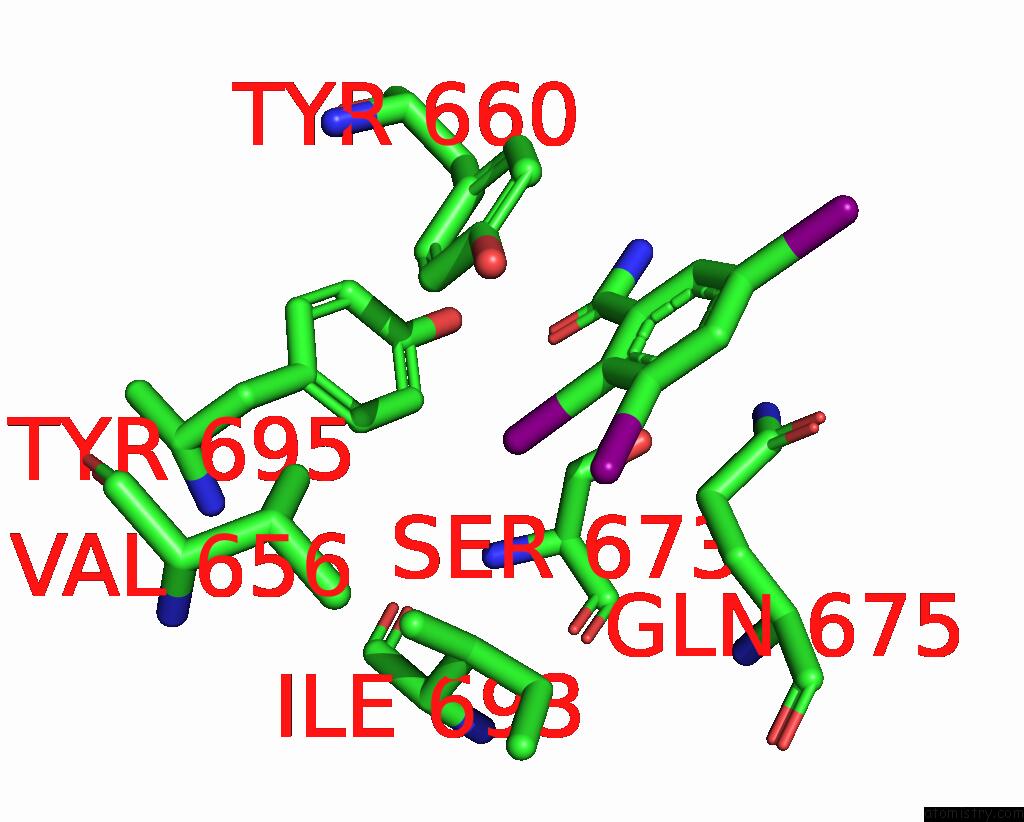
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 3 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Iodine binding site 4 out of 9 in 7qur
Go back to
Iodine binding site 4 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
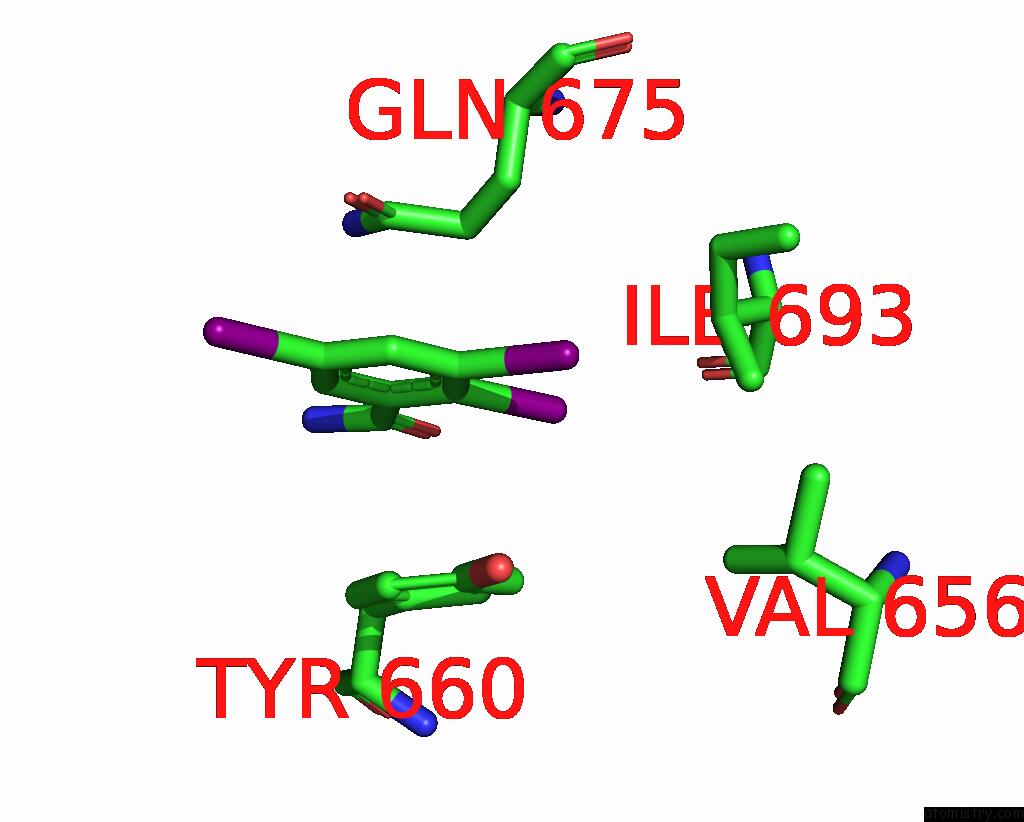
Mono view
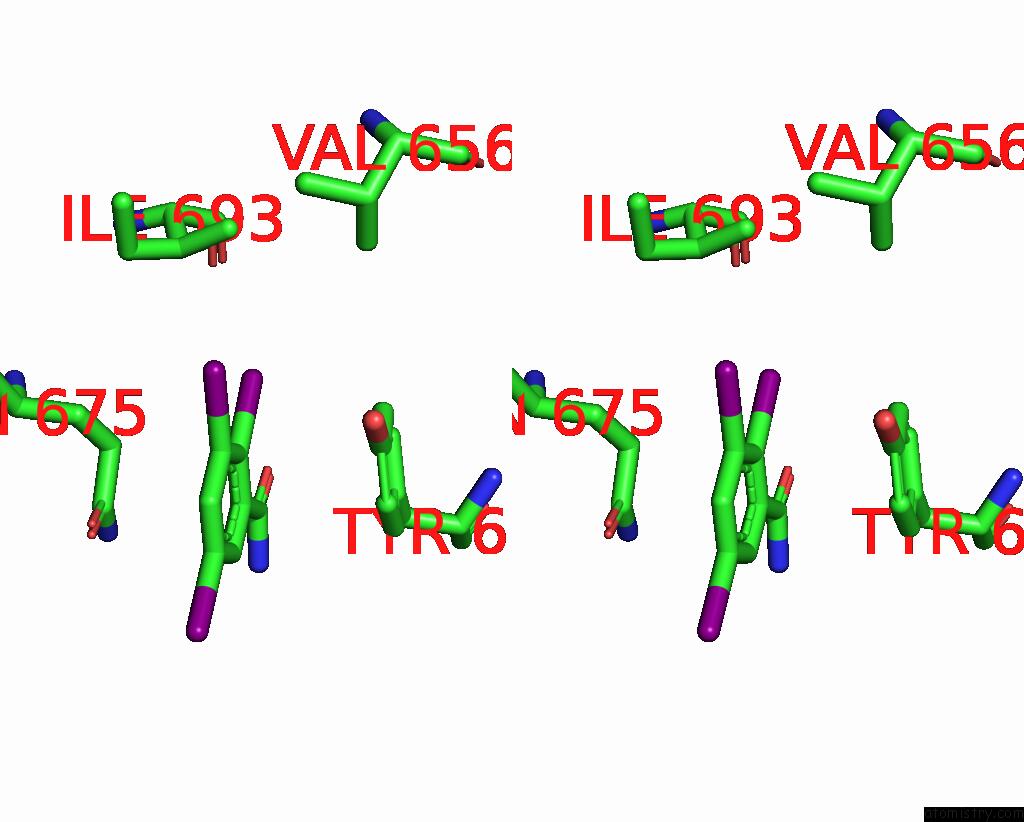
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 4 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Iodine binding site 5 out of 9 in 7qur
Go back to
Iodine binding site 5 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
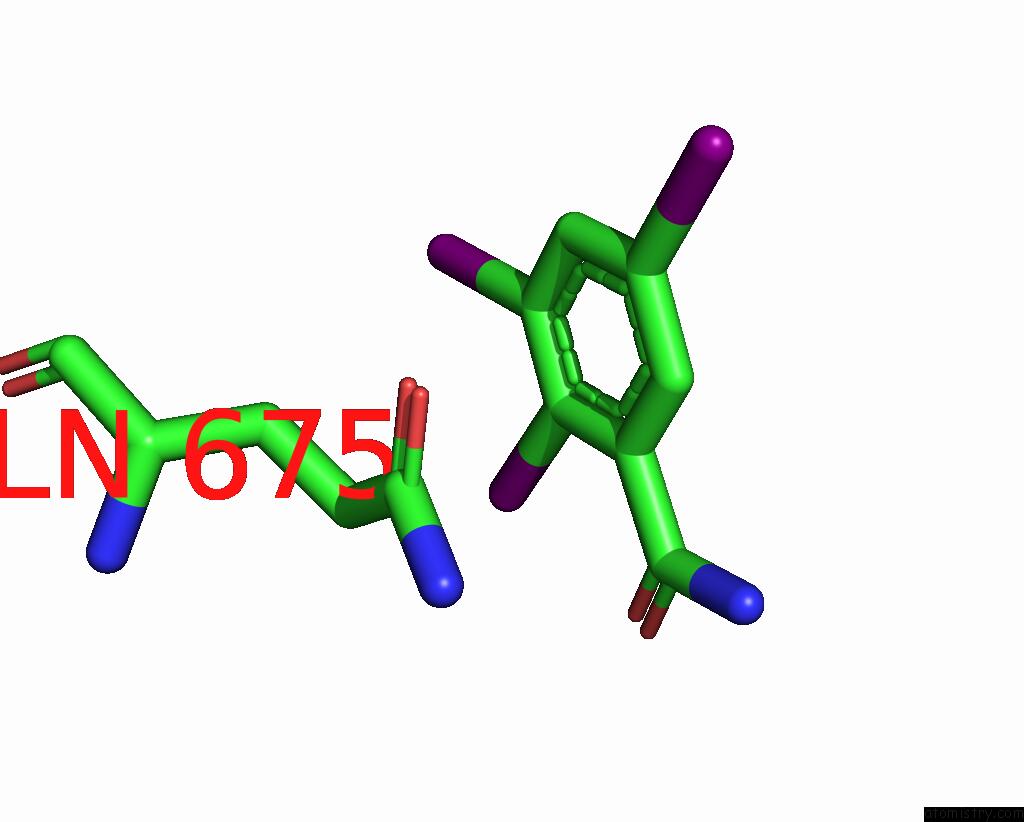
Mono view
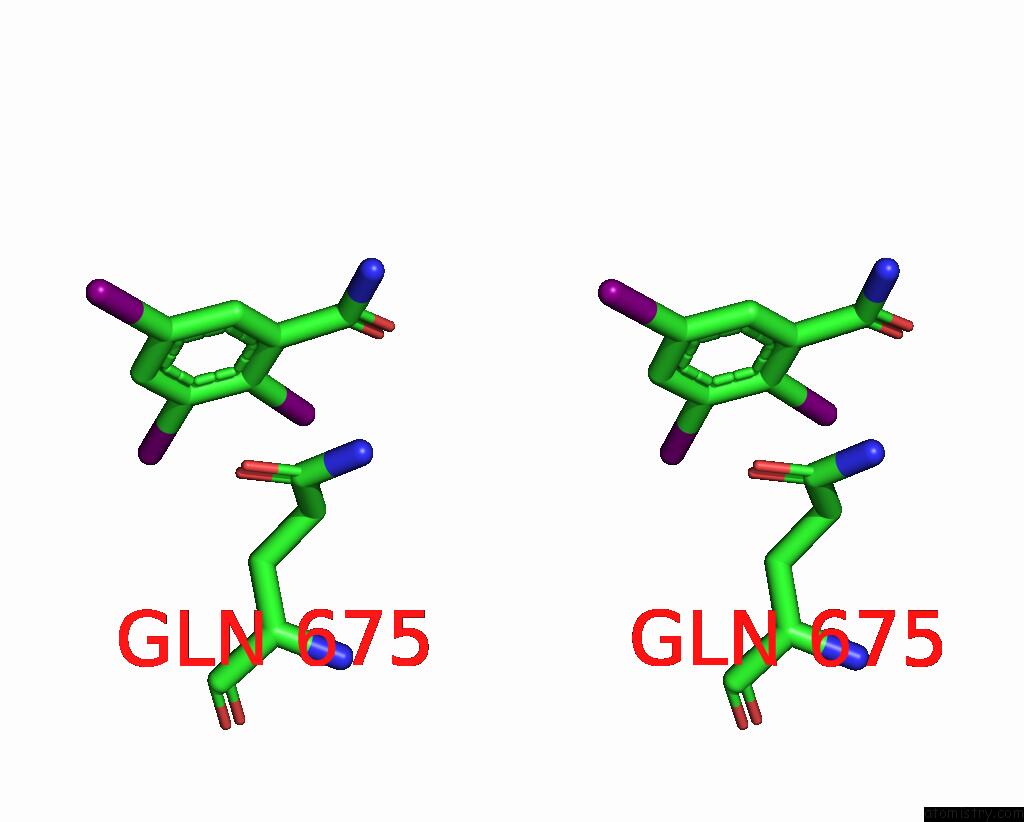
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 5 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Iodine binding site 6 out of 9 in 7qur
Go back to
Iodine binding site 6 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
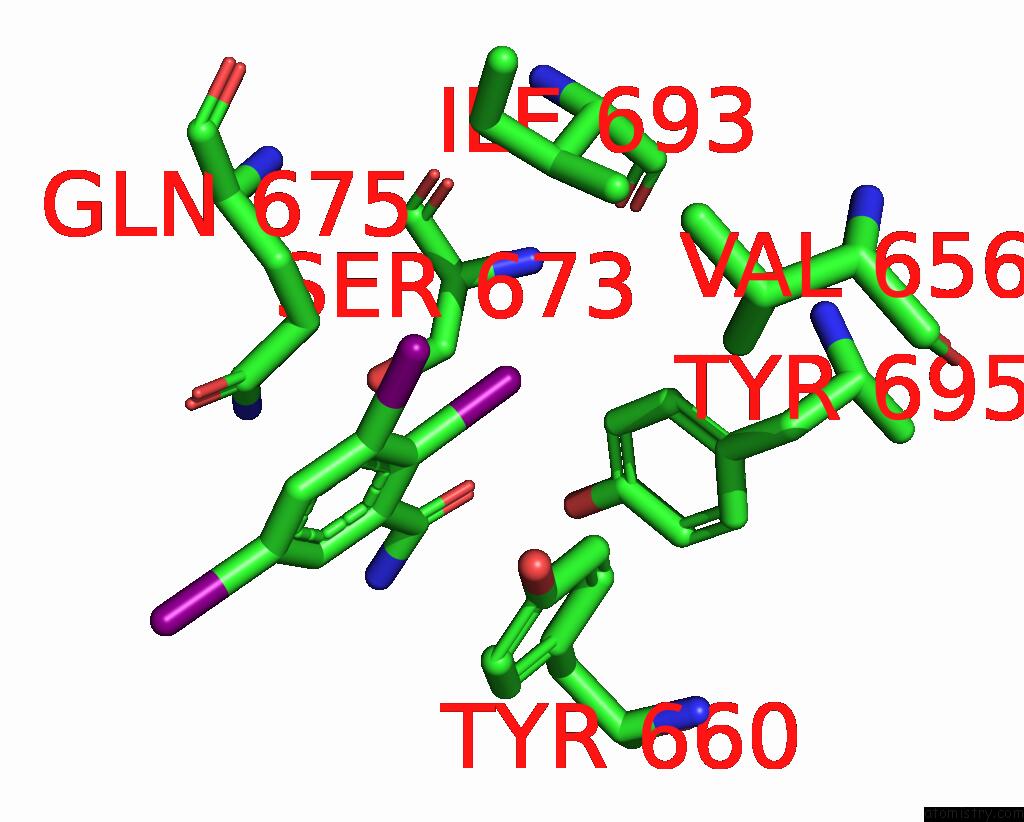
Mono view
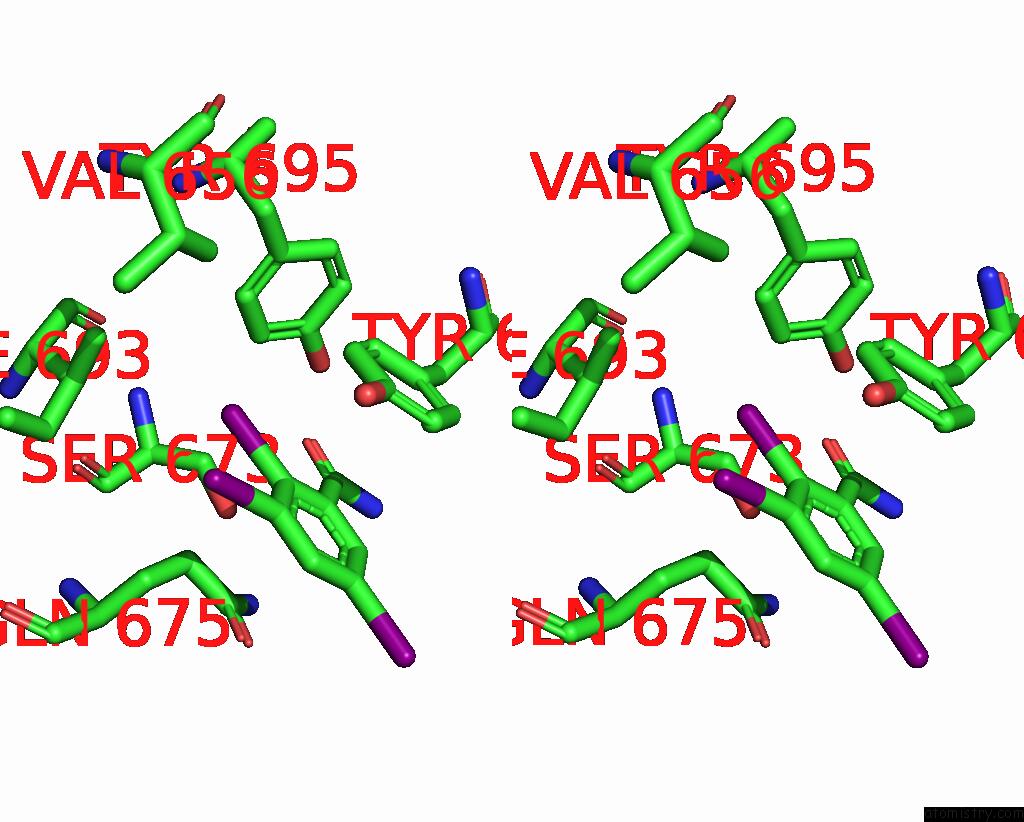
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 6 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Iodine binding site 7 out of 9 in 7qur
Go back to
Iodine binding site 7 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
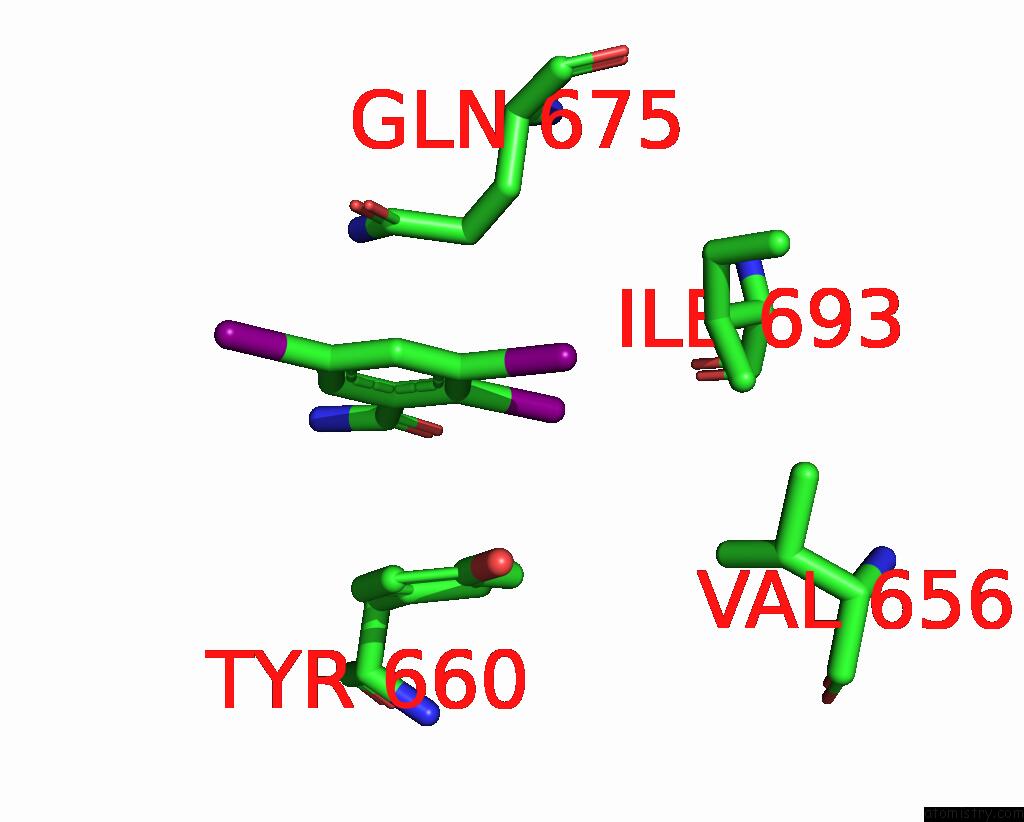
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 7 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Iodine binding site 8 out of 9 in 7qur
Go back to
Iodine binding site 8 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry
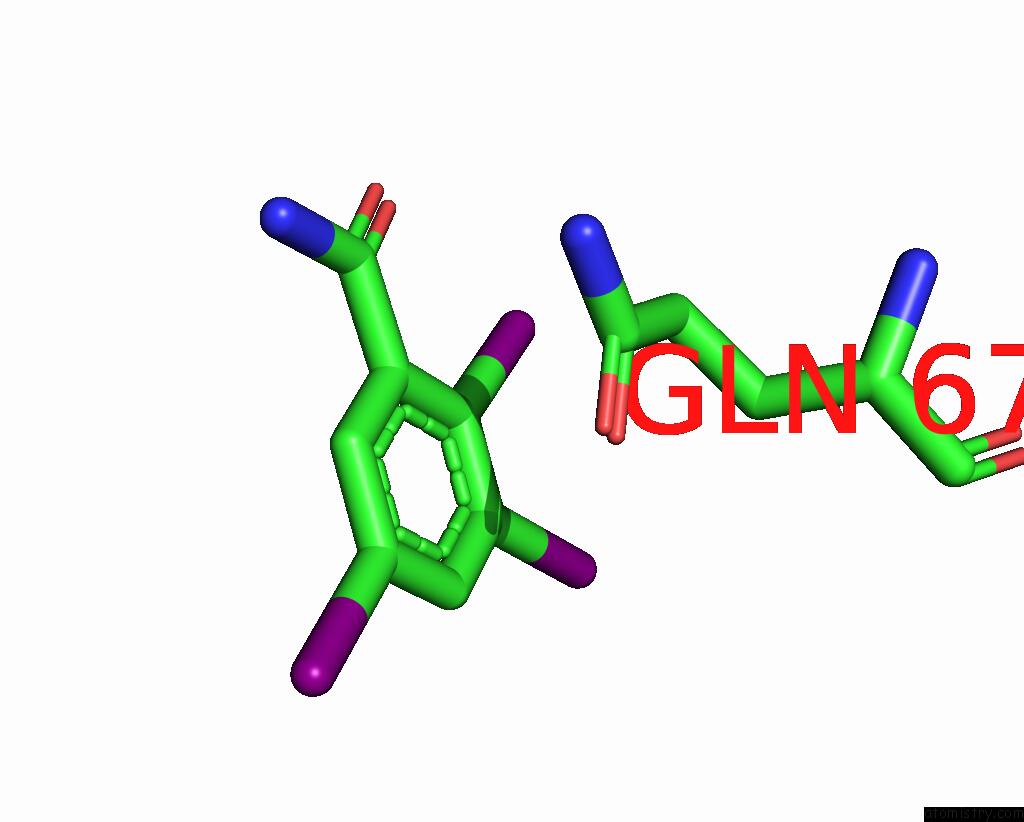
Mono view
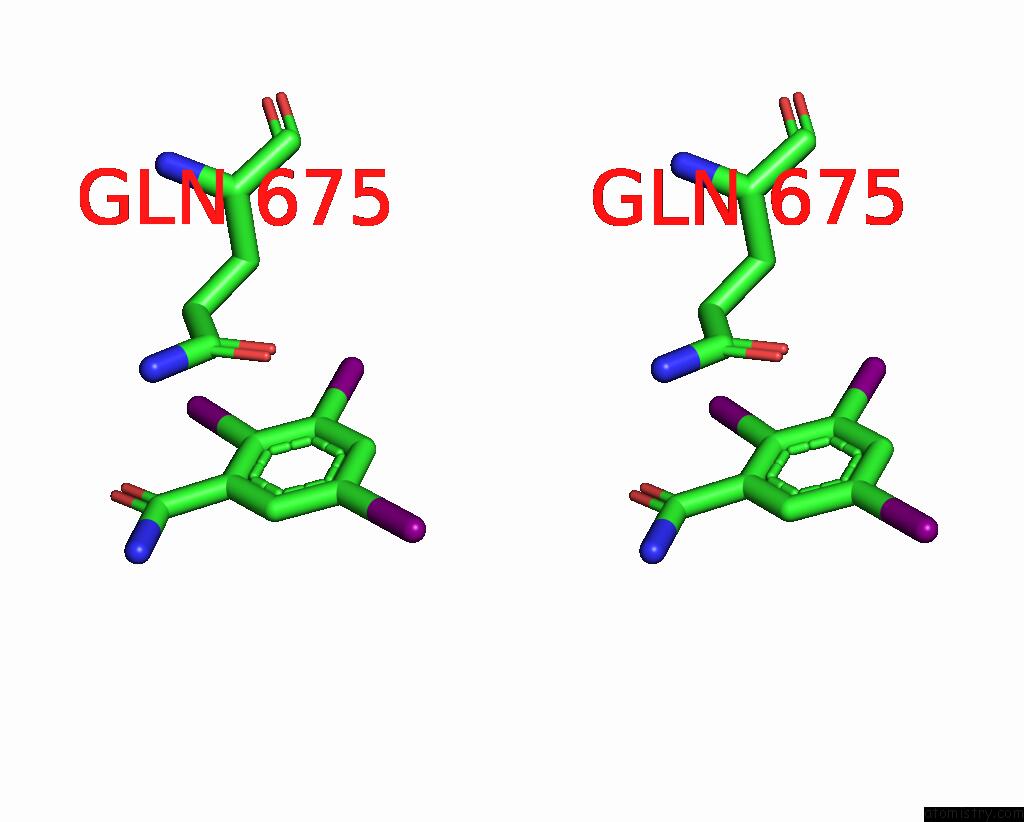
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 8 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Iodine binding site 9 out of 9 in 7qur
Go back to
Iodine binding site 9 out
of 9 in the Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry

Mono view
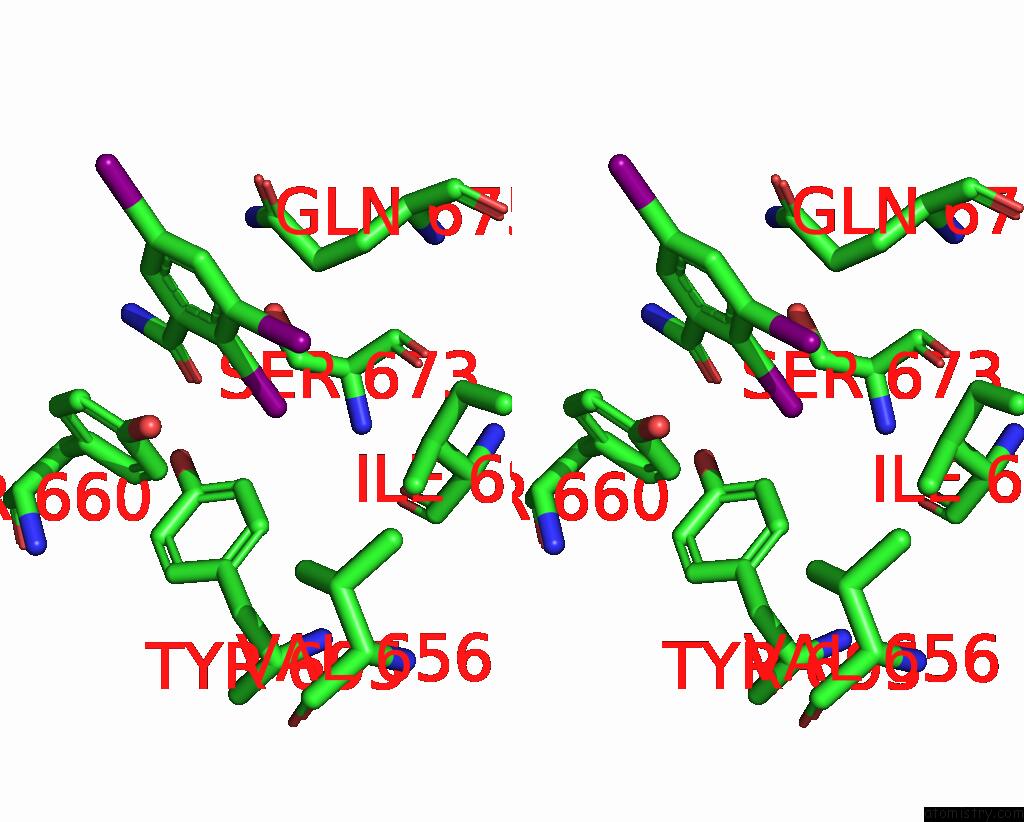
Stereo pair view

Mono view

Stereo pair view
A full contact list of Iodine with other atoms in the I binding
site number 9 of Sars-Cov-2 Spike with Ethylbenzamide-Tri-Iodo Siallyllactose, C3 Symmetry within 5.0Å range:
|
Reference:
C.J.Buchanan,
B.Gaunt,
P.J.Harrison,
Y.Yang,
J.Liu,
A.Khan,
A.M.Giltrap,
A.Le Bas,
P.N.Ward,
K.Gupta,
M.Dumoux,
T.K.Tan,
L.Schimaski,
S.Daga,
N.Picchiotti,
M.Baldassarri,
E.Benetti,
C.Fallerini,
F.Fava,
A.Giliberti,
P.I.Koukos,
M.J.Davy,
A.Lakshminarayanan,
X.Xue,
G.Papadakis,
L.P.Deimel,
V.Casablancas-Antras,
T.D.W.Claridge,
A.M.J.J.Bonvin,
Q.J.Sattentau,
S.Furini,
M.Gori,
J.Huo,
R.J.Owens,
C.Schaffitzel,
I.Berger,
A.Renieri,
J.H.Naismith,
A.J.Baldwin,
B.G.Davis.
Pathogen-Sugar Interactions Revealed By Universal Saturation Transfer Analysis. Science V. 377 M3125 2022.
ISSN: ESSN 1095-9203
PubMed: 35737812
DOI: 10.1126/SCIENCE.ABM3125
Page generated: Mon Aug 12 01:59:18 2024
ISSN: ESSN 1095-9203
PubMed: 35737812
DOI: 10.1126/SCIENCE.ABM3125
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF